Chi Lab
Research Interests
We are using functional genomic approaches to investigate the nutrient signaling and stress adaptations of cancer cells when exposed to various nutrient deprivations and microenvironmental stress conditions. Recently, we focus on two areas. First, we are elucidating the genetic determinants and disease relevance of ferroptosis, a newly recognized form of cell death. Second, we have identified the mammalian stringent response pathway which is highly similar to bacterial stringent response, but with some very interesting twists and novel mechanisms.
Research

A. The genetic determinants and disease relevance of ferroptosis
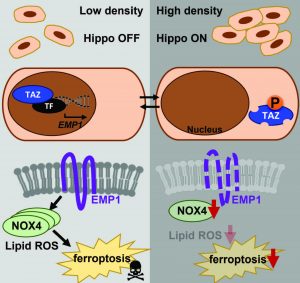 Ferroptosis is a newly recognized form of cell death that is characterized by iron dependency and lipid peroxidation. The importance of ferroptosis is being recognized in many human diseases, including cancers, ischemia injuries, and neurodegeneration. Previously, we have identified the profound cystine addiction of renal cell carcinoma (1), breast cancer cells (2, 3), and ovarian cancer cells (4). Based on the concept that cystine deprivation triggers the ferroptosis due to the unopposed oxidative stresses, we have performed functional genomic screens to identify many novel genetic determinants of ferroptosis. For example, we have found that DNA damage response and ATM kinase regulate ferroptosis via affecting iron metabolism (5). This finding supports the potential of ionizing radiation to trigger DNA damage response and synergize with ferroptosis to treat human cancers. In addition, we found that ferroptosis is highly regulated by cell density. When cells are grown at low density, they are highly susceptible to ferroptosis. In contrast, the same cells become resistant to ferroptosis when grown at high density and confluency. we have found the Hippo pathway effectors TAZ and YAP are responsible for the cell density-dependent ferroptosis (4, 6, 7). Right now, we are pursuing several other novel determinants of ferroptosis that will reveal surprising insights into this new form of cell death.
Ferroptosis is a newly recognized form of cell death that is characterized by iron dependency and lipid peroxidation. The importance of ferroptosis is being recognized in many human diseases, including cancers, ischemia injuries, and neurodegeneration. Previously, we have identified the profound cystine addiction of renal cell carcinoma (1), breast cancer cells (2, 3), and ovarian cancer cells (4). Based on the concept that cystine deprivation triggers the ferroptosis due to the unopposed oxidative stresses, we have performed functional genomic screens to identify many novel genetic determinants of ferroptosis. For example, we have found that DNA damage response and ATM kinase regulate ferroptosis via affecting iron metabolism (5). This finding supports the potential of ionizing radiation to trigger DNA damage response and synergize with ferroptosis to treat human cancers. In addition, we found that ferroptosis is highly regulated by cell density. When cells are grown at low density, they are highly susceptible to ferroptosis. In contrast, the same cells become resistant to ferroptosis when grown at high density and confluency. we have found the Hippo pathway effectors TAZ and YAP are responsible for the cell density-dependent ferroptosis (4, 6, 7). Right now, we are pursuing several other novel determinants of ferroptosis that will reveal surprising insights into this new form of cell death.
B. A new stress pathway – mammalian stress response
 All living organisms encounter a wide variety of nutrient deprivations and environmental stresses. Therefore, all organisms have developed various mechanisms to respond and promote survival under stress. In bacteria, the main strategy is “stringent response” triggered by the accumulation of the alarmone (p)ppGpp (shortened to ppGpp below) via regulation of its synthetase RelA and its hydrolase SpoT (8). The ppGpp binds to the transcription factor DksA and RNA polymerase to orchestrate extensive transcriptional changes that repress proliferation and promote stress survival (8, 9). While highly conserved among bacteria, the stringent response had not been reported in metazoans. However, a recent study identified Drosophila and human MESH1 (Metazoan SpoT Homolog 1) as the homologs of the ppGpp hydrolase domain of the bacterial SpoT (10). Both MESH1 proteins exhibit ppGpp hydrolase activity, and the deletion of Mesh1 in Drosophila led to a transcriptional response reminiscent of the bacterial stringent response (10). Recently, we have found that the genetic removal of MESH1 in tumor cells triggers extensive transcriptional changes and confers protection against oxidative stress-induced ferroptosis (11). Importantly, MESH1 removal also triggers proliferative arrest and other robust anti-tumor effects. Therefore, MESH1 knockdown leads to both stress survival and proliferation arrest, two cardinal features highly reminiscent of the bacterial stringent response. Therefore, we termed this pathway as “mammalian stringent response” (12). We have found that NADPH is the relevant MESH1 in the contexts of ferroptosis (13). Now, we are investigating how MESH1 removal leads to proliferation of arrests and anti-tumor phenotypes. Furthermore, we have found several other substrates of MESH1. We are investigating their function using culture cells, MESH1 KO mice, and other model organisms.
All living organisms encounter a wide variety of nutrient deprivations and environmental stresses. Therefore, all organisms have developed various mechanisms to respond and promote survival under stress. In bacteria, the main strategy is “stringent response” triggered by the accumulation of the alarmone (p)ppGpp (shortened to ppGpp below) via regulation of its synthetase RelA and its hydrolase SpoT (8). The ppGpp binds to the transcription factor DksA and RNA polymerase to orchestrate extensive transcriptional changes that repress proliferation and promote stress survival (8, 9). While highly conserved among bacteria, the stringent response had not been reported in metazoans. However, a recent study identified Drosophila and human MESH1 (Metazoan SpoT Homolog 1) as the homologs of the ppGpp hydrolase domain of the bacterial SpoT (10). Both MESH1 proteins exhibit ppGpp hydrolase activity, and the deletion of Mesh1 in Drosophila led to a transcriptional response reminiscent of the bacterial stringent response (10). Recently, we have found that the genetic removal of MESH1 in tumor cells triggers extensive transcriptional changes and confers protection against oxidative stress-induced ferroptosis (11). Importantly, MESH1 removal also triggers proliferative arrest and other robust anti-tumor effects. Therefore, MESH1 knockdown leads to both stress survival and proliferation arrest, two cardinal features highly reminiscent of the bacterial stringent response. Therefore, we termed this pathway as “mammalian stringent response” (12). We have found that NADPH is the relevant MESH1 in the contexts of ferroptosis (13). Now, we are investigating how MESH1 removal leads to proliferation of arrests and anti-tumor phenotypes. Furthermore, we have found several other substrates of MESH1. We are investigating their function using culture cells, MESH1 KO mice, and other model organisms.
C. Genomic and single cell RNA analysis of Red Blood Cells
Red blood cells (RBC) are responsible for oxygen delivery to muscles during vigorous exercise. Therefore, many doping efforts focus on increasing RBC number and function to boost athletic performance during competition. For many decades, RBC were thought to be merely identical “sacs of hemoglobin” with no discernable differences due to factors such as age or pre-transfusion storage time. Additionally, because RBC lose their nuclei during terminal differentiation, they were not believed to retain any genetic materials. These long-held beliefs have now been disproven and the results have significant implications for detecting autologous blood transfusion (ABT) doping in athletes. We were among the first to discover that RBCs contain abundant and diverse species of RNAs. Using this knowledge, we subsequently optimized protocols and performed genomic analysis of the RBC transcriptome in sickle cell disease; these results revealed that heterogeneous RBCs could be divided into several subpopulations, which had implications for the mechanisms of malaria resistance. As an extension of these studies, we used high resolution Illumina RNA-Seq approaches to identify hundreds of additional known and novel microRNAs, mRNAs, and other RNA species in RBCs. This dynamic RBC transcriptome represents a significant opportunity to assess the impact that environmental factors (such as pre-transfusion refrigerate storage) on the RBC transcriptome. We have now identified a >10-fold change in miR-720 as well as several other RNA transcripts whose levels are significantly altered by RBC storage (14) which gained significant press coverage. We are pursuing the genomic and single cell analysis of RNA transcriptome in the context of blood doping, sickle cell diseases and other red cell diseases.

- Tang X, Wu J, Ding CK, Lu M, Keenan MM, Lin CC, et al. Cystine Deprivation Triggers Programmed Necrosis in VHL-Deficient Renal Cell Carcinomas. Cancer Res. 2016;76(7):1892-903.
- Tang X, Ding CK, Wu J, Sjol J, Wardell S, Spasojevic I, et al. Cystine addiction of triple-negative breast cancer associated with EMT augmented death signaling. Oncogene. 2017;36(30):4379.
- Lin CC, Mabe NW, Lin YT, Yang WH, Tang X, Hong L, et al. RIPK3 upregulation confers robust proliferation and collateral cystine-dependence on breast cancer recurrence. Cell Death Differ. 2020.
- Yang WH, Huang Z, Wu J, Ding C-KC, Murphy SK, Chi J-T. A TAZ-ANGPTL4-NOX2 axis regulates ferroptotic cell death and chemoresistance in epithelial ovarian cancer. Molecular Cancer Research. 2019: molcanres.0691.2019.
- Chen PH, Wu J, Ding CC, Lin CC, Pan S, Bossa N, et al. Kinome screen of ferroptosis reveals a novel role of ATM in regulating iron metabolism. Cell Death Differ. 2019.
- Yang W-H, Chi J-T. Hippo pathway effectors YAP/TAZ as novel determinants of ferroptosis. Molecular & Cellular Oncology. 2019:1699375.
- Yang WH, Ding CKC, Sun T, Hsu DS, Chi JT. The Hippo Pathway Effector TAZ Regulates Ferroptosis in Renal Cell Carcinoma Cell Reports. 2019;28(10):2501-8.e4.
- Potrykus K, Cashel M. (p)ppGpp: still magical? Annu Rev Microbiol. 2008;62:35-51.
- Kriel A, Bittner AN, Kim SH, Liu K, Tehranchi AK, Zou WY, et al. Direct regulation of GTP homeostasis by (p)ppGpp: a critical component of viability and stress resistance. Mol Cell. 2012;48(2):231-41.
- Sun D, Lee G, Lee JH, Kim HY, Rhee HW, Park SY, et al. A metazoan ortholog of SpoT hydrolyzes ppGpp and functions in starvation responses. Nat Struct Mol Biol. 2010;17(10):1188-94.
- Dixon SJ, Lemberg KM, Lamprecht MR, Skouta R, Zaitsev EM, Gleason CE, et al. Ferroptosis: an iron-dependent form of nonapoptotic cell death. Cell. 2012;149(5):1060-72.
- Ding C-KC, Rose J, Wu J, Sun T, Chen K-Y, Chen P-H, et al. Mammalian stringent-like response mediated by the cytosolic NADPH phosphatase MESH1. bioRxiv. 2018.
- Ding C-KC, Rose J, Sun T, Wu J, Chen P-H, Lin C-C, et al. MESH1 is a cytosolic NADPH phosphatase that regulates ferroptosis. Nature Metabolism. 2020.
- Yang WH, Doss JF, Walzer KA, McNulty SM, Wu J, Roback JD, et al. Angiogenin-mediated tRNA cleavage as a novel feature of stored red blood cells. Br J Haematol. 2018.
Biography
Jen-Tsan Ashley Chi, MD PhD is an associate professor in the Department of Molecular Genetics and Microbiology and Center for Genomic and Computational Biology. He received his MD from National Taiwan University and his PhD from Stanford University. After his PhD, he was a post-doctoral fellow in Dr. Patrick Brown’s lab in Stanford University focusing on using cDNA microarrays to elucidate the cellular differentiation as well as human pathogenesis. He joined the faculty at MGM and IGSP in 2004.
Originally from Taiwan, Dr. Chi originally received his MD from National Taiwan University. Shortly afterwards, he received his PhD and completed postdoctoral training with Dr. Patrick Brown at Stanford University. A stellar physician-scientist, he joined the faculty here at Duke as an assistant professor in the Department of Molecular Genetics and Microbiology working closely within the Institute of Genome Sciences and Policy. Dr. Chi has conducted microarray research and published many studies on the use of genomic analysis to dissect the influence of hypoxia and various tumor microenvironmental stresses on cancer cells to dissect the heterogeneity of human cancers. His research program has been focused on the use gene signatures (hypoxia, lactic acidosis) obtained in a tissue culture setting to annotate and analyze tumor phenotypes and other characteristics of human cancers.
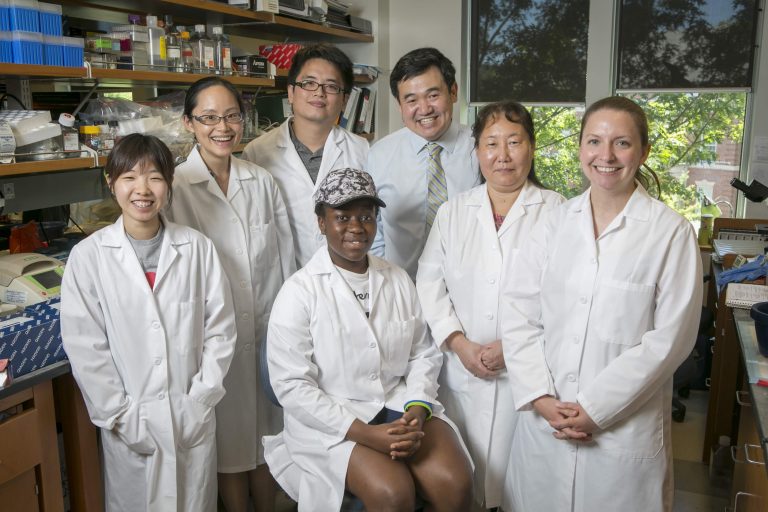
Lab Members
Publications
Representative Publications
Chang, H.Y., Chi, J.T., Dudoit, S., Bondre, C., van de Rijn, M., Botstein, D. and Brown, P. O. (2002) Diversity, topographic differentiation, and positional memory in human fibroblasts. Proceeding of National Academy of Science USA 99:12877-12882.
Chi, J.T., Chang, H.Y., Wang, N.N., Chang, D.S., Dunphy, N. and Brown, P.O. (2003) Genomewide view of gene silencing by small interfering RNAs. Proceeding of National Academy of Science USA 100:6343-6346.
Chi, J.T., Chang, H.Y., Haraldsen, G., Jahnsen, F.L., Troyanskaya, O.G., Chang, D.S., Wang, Z., Rockson, S.G., van de Rijn, M., Botstein, D. and Brown, P.O. (2003) Endothelial cell diversity revealed by global expression profiling. Proceeding of National Academy of Science USA 100:10623-10628.
Whitfield, M.L., Finlay, D., Murray, J.I., Troyanskaya, O.,Chi, J.T., Brown, P.O., Botstein, D. and Connolly, M.K. (2003) Patterns of gene expression in scleroderma. Proceeding of National Academy of Science USA100:12319-12324.
Chang, H.Y., Sneddon, J.B., Alizadeh, A.A., Sood R., West, R.B., Montgomery, K., Chi, J. T., van de Rijn, M., Botstein, D. and Brown, P.O. (2004) Gene expression signature of fibroblast serum response predicts human cancer progression–similarities between tumors and wounds. PLoS Biology 2:e7.
Chi, J.T., Wang, Z., Nuyten, D.S.A., Rodriguez, E.H., Schaner, M.E., Salim, A., Wang, Y., Kristensen, G.B., Helland, A., Børresen-Dale, A., Giaccia, A., Longaker, M.T., Hastie, T., Yang, Y.P., van de Vijver, M.J. and Brown, P.O. (2006) Gene expression programs in response to hypoxia; cell type specificity and prognostic significance in human cancers. PLoS Medicine 3:e47.
Erler, J.T., Bennewith, K.L., Nicolau, M., Dornhofer, N., Kong, C., Le, Q.T., Chi, J.T., Jeffrey, S.S., Giaccia, A.J. (2006) Lysyl oxidase is essential for hypoxia-induced metastasis. Nature 440:1222-1226.
Myers, J.W., Chi, J.T., Gong, D, Schaner, M.E., Brown, P.O., Ferrell, J.E. (2006) Minimizing off target effects by using diced siRNAs for RNA interference. Journal of RNAi and Gene Silencing 2:181-194.
Moon, E.J., Brizel, D.M., Chi, J.T. and Dewhirst, M.W. (2007) The potential role of intrinsic hypoxia markers as prognostic variables in cancer. Antioxidants & Redox Signaling 9:1237-1294.
Chi, J.T., Rodriguez, E.H., Wang, Z., Nuyten, D., Mukherjee, S., van de Rijn, M., Vijver, M.J., Hastie, T. and Brown, P. O. (2007) Gene expression programs of human smooth muscle cells: Tissue-specific differentiation and prognostic significance in breast cancers. PLoS Genetics 3:e164.
Gottwein, E., Mukherjee, N., Sachse, C., Frenzel, C., Majoros, W.H., Chi, J.T., Braich, R., Manoharan, M., Soutschek, J., Ohler, U. and Cullen, B. (2007) A viral microRNA functions as an ortholog of cellular miR-155. Nature 450:1096-1099.
Edelman, E., Guinney, J., Chi, J.T., Febbo, P.G., Mukherjee, S. (2008) Modeling cancer progression via pathway dependencies. PLoS Computational Biology 4:e28.
Chen, S.Y., Wang, Y., Telen, M.J., Chi, J.T. (2008) The genomic analysis of erythrocyte microRNAs expression in sickle cell diseases. PLoS ONE 3:e2360.
Nuyten, D.S., Hastie, T., Chi, J.T., Chang, H.Y., Vijver, M.J. (2008) Combining biological gene expression signatures in predicting outcome in breast cancer: An alternative to supervised classification. European Journal of Cancer 44: 2319-2329.
Chen, J.L., Lucase, J.E., Schroede,r T., Mori, S., Nevins, J.R., Dewhirst, M.W., West, M., Chi, J.T. (2008) Genomic analysis of lactic acidosis and acidosis response in human cancers. PLoS Genetics 4:e1000293.
Lucas, J.E., Carvalho, C.M., Chen, J.L., Chi, J.T. and West, M. (2009) Cross-study projections of Genomic biomarkers: An evaluation in cancer genomics. PLoS One 4:e4523.
Chan, A.S., Kawahara, T.L., Sutphin, P.D., Change, H.Y., Chi, J.T., Giaccia, A. (2009) Tumor vasculature is regulated by PHD2- mediated angiogenesis and bone marrow-derived cell recruitment. Cancer Cell 5:527-538.
Merl, D., Chen, J.L., Chi, J.T. and West, M. (2009) An integrative analysis of cancer gene expression studies using Bayesian latent factor modeling. Annals of Applied Statistics 3:1675-1694.
Pogozelski, A.R., Geng, T., Li, P., Yin, X., Lira, V.A., Zhang, M., Chi, J.T., and Yan, Z. (2009) p38gamma mitogen-activated protein kinase is a key regulator in skeletal muscle metabolic adaptation in mice. PLoS One 4:e7934.
Himburg, H.A. Muramoto, G.G., Daher, P., Meadows, S.K., Russell, J.L., Doan, P., Chi, J.T., Salter, A.B., Lento, W.E., Reya, T., Chao, N.J., and Chute, J.P. (2010) Pleiotrophin regulates the expansion and regeneration of hematopoietic stem cells. Nature Medicine 16:475-482.
Lindsay, R.G. and Chi, J.T. (2010) Contact lens management of infantile aphakia. Clinical and Experimental Optometry 93:3-14. Review
Chen, J.L, Merl, D., Peterson, C., Wu, J, Liu, P, Ayer, D, West M, and Chi, J.T. (2010) Lactic acidosis triggers starvation response with paradoxical induction of TXNIP through MondoA. PLOS Genetics 6:e1001093.
Sangokoya, C., Telen, M. J., Chi, J. T. (2010) MicroRNA miR-144 modulates oxidative stress tolerance and associates with anemia severity in sickle cell disease. Blood 116: 4338-4348.
Lucas J.E., Kung H.N., Chi J.T. (2010) Latent factor analysis to discover pathway-associated putative segmental aneuploidies in human cancers. PLoS Computational Biology 6:e1000920.
Sangokoya C., Lamonte G., Chi J.T. (2010) Isolation and Characterization of MicroRNAs of Human Mature Erythrocytes Methods. Molecular Biology 667:193-203.
Chi J.T. #, Thrall D.E., Jiang C., Snyder S., Fels D., Landon C., McCall L., Lan L., Hauck M., MacFall J.R., Viglianti B.L. and Dewhirst M.W. Comparison of genomics and functional imaging from canine sarcomas treated with thermoradiotherapy predicts therapeutic response and identifies combination therapeutics. Clinical Cancer Research 17:2549-60, 2011 PMCID: PMC3078971 #Corresponding author.
Gatza M.L., Kung H.N., Blackwell K.L., Dewhirst M.W., Marks J.R., Chi J.T. Analysis of tumor environmental response and oncogenic pathway activation identifies distinct basal and luminal features in HER2-related breast tumor subtypes. Breast Cancer Research 13: R62. 2011 “Highly Accessed” PMCID: PMC3218951
Lacsina J.R., LaMonte G., Nicchitta, C.V., and Chi J.T. Polysome profiling of the malaria parasite Plasmodium falciparum. Molecular & Biochemical Parasitology 179:42-6. Epub 2011 May 13. PMCID: PMC3135974
Akslen L.A., Straume O., Geisler S., Sorlie T., Chi J.T., Aas T., Børresen-Dale A.L., Lønning P.E. Glomeruloid microvascular proliferation is associated with lack of response to chemotherapy in breast cancer. British Journal of Cancer 105:9-12, 2011. PMCID: PMC3137417
Lewis D.A., Stashenko G.J., Akay O.M., Price L.I., Owzar K., Ginsburg G.S., Chi J.T., and Ortel T.L. Whole blood gene expression analyses in patients with single versus recurrent venous thromboembolism. Thrombosis Research 128:536-40, 2011.
Chan D.A., Sutphin P.D., Nguyen P., Turcotte S., Lai E.W., Banh A., Reynolds G.E., Chi J.T., Wu J., Solow-Cordero D.E., Bonnet M., Flanagan J.U., Bouley D.M., Graves E.E., Denny W.A., Hay M.P., and Giaccia A.J. Targeting GLUT1 and the Warburg effect in renal cell carcinoma by chemical synthetic lethality. Scientific Translational Medicine 3:94ra70, 2011.
Kung H.N., Marks J.R., and Chi J.T. Glutamine synthetase is a genetic determinant of cell type-specific glutamine independence in breast epithelia. PLoS Genetics 7(8):e1002229, 2011. PMCID: PMC3154963.




