Research
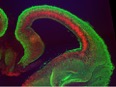
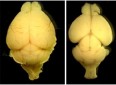
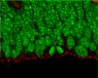
Our laboratory studies embryonic brain development, with a specific focus on neurogenesis of the cerebral cortex. During neurogenesis, neural progenitor populations produce neurons. Precise control of progenitor divisions during development helps dictate the size, structure, and function of the adult brain. Aberrant neurogenesis can result in several neurodevelopmental disorders such as microcephaly (reduced brain size associated with intellectual disability) and autism spectrum disorder. The mechanisms that control neurogenesis remain poorly understood, which limits our ability to understand the etiology of these disorders. Our goal is to help fill this void by uncovering genetic and cellular regulation of neurogenesis.
A major research direction of our lab is to define genetic mechanisms of brain size, with a focus on post-transcriptional regulation. We study requirements for RNA binding proteins and for post-transcriptional regulation in dynamic neural progenitor behavior and function. The RNA binding proteins studied are also associated with neurodevelopmental pathology including brain malformations. A second focus of our research is to understand how genetic noncoding loci (termed human accelerated enhancers) contribute to unique features of the human brain. We study enhancers relevant for both brain size and neural progenitor proliferation.
The lab employs a repertoire of genetic and cell biological tools including mouse genetics, ex vivo and in vitro live imaging, and genomics. Using multidisciplinary approaches helps give us mechanistic insights at molecular, cellular, and organismal levels. Our long-term objective is to help broaden our fundamental understanding of how the brain is built, how stem cells behave, and the etiology of developmental diseases.
Please see our lab website (https://sites.duke.edu/silverlab) for more detailed information on research projects in the lab.
Biography
Debby Silver received her B.S. in Biology from Tufts University in 1993. Following her undergraduate degree she worked with Dr. James Sellers at the NIH (National Heart Lung and Blood Institute), where she applied biochemistry to study the cytoskeleton. Her graduate training was with Dr. Denise Montell of The Johns Hopkins University School of Medicine. For her thesis, she discovered a novel essential role for the JAK/STAT signaling complex in cell migration of Drosophila ovarian epithelial cells and human ovarian cancer cells. This work was recognized with several graduate student awards, including the national Weintraub award.
For her postdoctoral studies, Dr. Silver trained with Dr. William Pavan of the National Human Genome Research Institute, where she used mouse genetics to study neural development in mice, with a focus on regulation of neural crest and cortical development. This work led to the discovery of an RNA binding complex essential for cerebral cortex development and brain size. Her research was funded both extramurally and intramurally (NIGMS PRAT fellowship (Pharmacology Research and Training) and NINDS NIH Pathway to Independence Award (K99/R00).
In 2010, Dr. Silver joined the MGM Department at Duke University Medical Center and the Duke Institute for Brain Sciences. In 2011 she became a secondary faculty member in the Department of Neurobiology and the Department of Cell Biology. Her lab studies embryonic brain development, with a focus on the process of neurogenesis in which neurons are produced from neural progenitors. The lab aims to better understand the genetic and cellular mechanisms controlling neurogenesis and contributing to neurodevelopmental disorders such as microcephaly and autism. A major focus of the lab is on post-transcriptional regulation of neural stem cells. The lab is also interested in elucidating genomic loci that are relevant for neurogenesis and evolution of the human neocortex.
Lab Members
Publications
Top 10 Publications:
- Miller, EE+, Kobayashi, GS+, Musso, CM, Allen, M, Ishiy, FAA, Caires Junior, LC, Guimarães, ESG, Griesi-Oliveira, K, Zechi-Ceide, RM, Richieri-Costa, A, Bertola, DR, Passos-Bueno, MR*, Silver, DL*. EIF4A3 deficient human iPSCs and mouse models demonstrate neural crest defects that underlie Richieri-Costa-Pereira Syndrome. Human Molecular Genetics. June 15; 26 (12): 2177-2191. 2017, *co-corresponding authors, + co-first authors
Pilaz, L-J. and Silver, DL. Moving messages in the developing brain: Emerging roles for mRNA transport and local translation in neural stem cells.FEBs Letters. June; 591 (11): 1526-2539. 2017 Invited review article.
Lennox, AL, Mao, H, and Silver, DL. RNA on the brain: emerging layers of post-transcriptional regulation in cerebral cortex development. Wiley Interdisciplinary Reviews Developmental Biology. Aug 24 online. 2017 Invited review article.
Mitchell C, Silver DL. Enhancing our brains: Genomic mechanisms underlying cortical evolution. Invited review article. Semin Cell Dev Biol. Aug 29. pii: S1084-9521(17)30235-5. 2017
Pilaz, L-J, Lennox, A.L., Rouanet, J.P., and Silver, DL. Dynamic mRNA transport and local translation in radial glial progenitors of the developing brain. Current Biology. Online Nov. 30. pii: S0960-9822(16)31266-0. PMID: 27916527. 26: 1-10.
- Mao, H*, McMahon, JJ*, Tsai, Y-H., Wang, Z, and Silver, DL. Haploinsufficiency for core exon junction complex components disrupts neurogenesis and causes p53-mediated microcephaly. PLOS Genetics. Sept. 12.; 12(9): e1006282. 2016 * co-first authors.
Mao, H, Brown, H, and Silver, DL. Mouse models of Casc3 reveal developmental functions distinct from other components of the exon junction complex. RNA. Online Oct. 25. pii: rna.058826.116. PMID: 27780844.
Pilaz, LJ*, McMahon, JJ*, Miller, EE, Lennox, AL, Suzuki, A, Salmon, E, Silver, DL, Prolonged mitosis of neural progenitors alters cell fate in the developing brain. Jan 6; 89(1):83-99. 2016. *co-first authors.
Mao, H, Pilaz, LJ, McMahon, J, Golzio, C, Wu, D, Shi, L, Katsanis, N, and Silver, DL Rbm8a haploinsufficiency disrupts embryonic cortical development resulting in microcephaly. Journal of Neuroscience. May 6; 35(18): 7003-7018. (Featured on cover and in News and Views in same issue).
Boyd, JL, Skove, SL, Rouanet, JP, Pilaz, L-J, Bepler, T, Gordân, R, Wray, GA, and Silver, DL. Human-chimpanzee differences in a FZD8 enhancer alter cell-cycle dynamics in the developing neocortex. Current Biology. Mar 16; 25 (6) 772-9. 2015. (Featured in over 75 online and print forums including Science, National Geographic, NPR.)
To view a complete list of publications, click here.





